MBI Videos
Marc Suchard
-
 Marc SuchardPanel discussion led by Marc Suchard
Marc SuchardPanel discussion led by Marc Suchard -
 Marc Suchard
Marc SuchardTBD
-
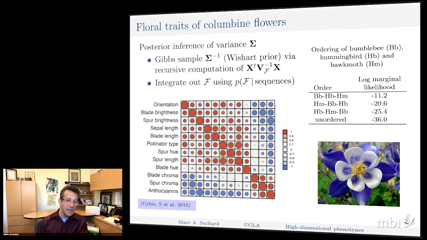
 Marc SuchardMassive numerical integration plagues the statistical inference of partially observed stochastic processes. An important biological example entertains partially observed continuous-time Markov chains (CTMCs) to model molecular sequence evolution. Joint inference of phylogenetic trees and codon-based substitution models of sequence evolution remains computationally impractical. Parallelizing data likelihood calculations is an obvious strategy; however, across a cluster-computer, this scales with the total number of processing cores, incurring considerable cost to achieve reasonable run-time.
Marc SuchardMassive numerical integration plagues the statistical inference of partially observed stochastic processes. An important biological example entertains partially observed continuous-time Markov chains (CTMCs) to model molecular sequence evolution. Joint inference of phylogenetic trees and codon-based substitution models of sequence evolution remains computationally impractical. Parallelizing data likelihood calculations is an obvious strategy; however, across a cluster-computer, this scales with the total number of processing cores, incurring considerable cost to achieve reasonable run-time.
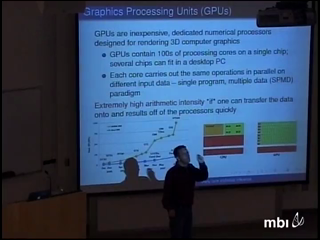
To solve this problem, I describe many-core computing algorithms that harness inexpensive graphics processing units (GPUs) for calculation of the likelihood under CTMC models of evolution. High-end GPUs containing hundreds of cores and are low-cost. These novel algorithms are particularly efficient for large state-spaces, including codon models, and large data sets, such as full genome alignments where we demonstrate up to 150-fold speed-up. I conclude with a discussion of the future of many-core computing in statistics and touch upon recent experiences with massively large and high-dimensional mixture models.